The UNC Flow Cytometry Core Facility (FlowCore) provides state-of-the-art flow cytometry services to the entire UNC-CH research community as well as to others in the Research Triangle Park area. A skilled staff provides help with experimental design, sample acquisition, data analysis, and sorting. Training is available to enable investigators and their staff to run the analyzers independently at reduced cost. A major part of our mission is to teach this technology to investigators, students, and staff. We are committed to providing guidance for the highest level of rigor and reproducibility in all aspects of experimental design, data acquisition, analysis and reporting. Please do not hesitate to contact us if you have any questions about flow cytometry, if you want to know if you can use it in your research, how to design experiments, prepare samples, or how to analyze your data.
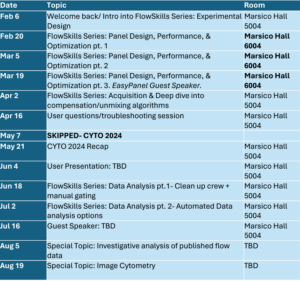
(Tentative upcoming events, check Teams Page for more details)
The core is located in the basement of the Marsico Hall, 125 Mason Farm Rd, Chapel Hill, NC. (Directions)
Follow us on Twitter: @UNC_Flow
Acknowledging the Core in Publications